May 19, 2021
Complete Omics today highlighted a decade-long scientific journey in clinical proteomics, marking ten years since the landmark 2011 publication in Proceedings of the National Academy of Sciences (PNAS) that first demonstrated the targeted detection of mutant proteins in body fluids. Over the past ten years, this work has evolved from detecting a small set of cancer-derived mutant proteins to building platforms capable of measuring thousands of proteins in human plasma, establishing a new foundation for molecular diagnostics and precision medicine.
2011: From 6 mutant protein biomarkers to a new realization: the plasma proteome is measurable and quantifiable
The origin of our effort in developing the ultimate clinical proteomics pipeline traces back to a landmark study demonstrating that mutant proteins produced by cancer driver mutations could be directly detected in body fluids using targeted mass spectrometry.
The study, “Mutant proteins as cancer-specific biomarkers.” Proc Natl Acad Sci U S A . 2011 Feb 8;108(6):2444-9. doi: 10.1073/pnas.1019203108. Epub 2011 Jan 19. showed that approximately 6 mutant peptide biomarkers could be simultaneously detected and quantified, representing one of the first demonstrations that genomic mutations could be translated into measurable mutant proteins detectable in complex biological fluids.
For the researchers who would later found Complete Omics, this discovery triggered an important realization:
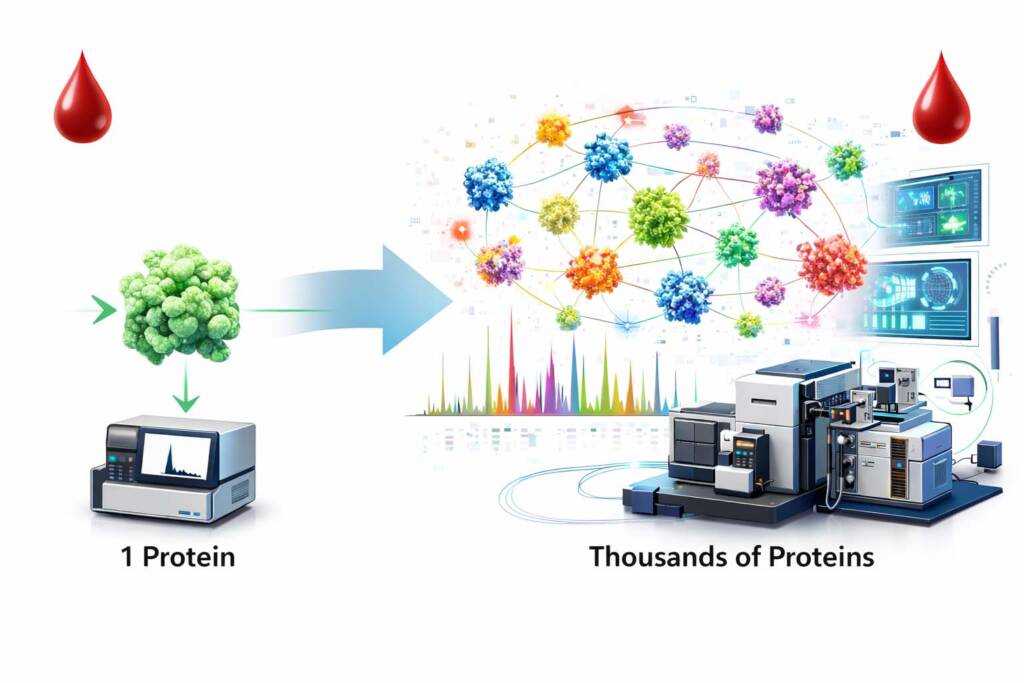
If a panel of mutant proteins could be reproducibly detected in plasma, then it should be possible to detect many more proteins from body fluids with the right analytical engineering and measurement strategies. This insight initiated a long-term effort to expand the measurable portion of the human proteome in complex biological samples.
2017: Expanding multiplex protein detection with SAFE-SRM
Building on this concept, the team significantly expanded the scale of multiplex protein detection.
In 2017, the group reported a major advance in targeted proteomics named the “SAFE-SRM” strategy, enabling the quantitative measurement of 652 protein biomarkers simultaneously for ovarian cancer screening applications. This work was reported in “Selected reaction monitoring approach for validating peptide biomarkers” Proc Natl Acad Sci U S A. 2017 Dec 19;114(51):13519-13524. The SAFE-SRM platform demonstrated that large multiplex panels of proteins could be quantitatively measured with high specificity, sensitivity, and reproducibility, addressing a long-standing challenge in plasma proteomics where the extreme dynamic range of blood proteins had historically limited analytical depth.
This milestone reinforced a growing hypothesis — the plasma proteome could become a scalable diagnostic medium if reproducible measurement technologies were systematically engineered.
After the ovarian cancer work, it became clear to us that, with sufficient effort, comprehensive detection of essentially all human proteins from body fluid samples is achievable. Through this work, we also accumulated substantial first-hand knowledge of the critical factors that must be considered to turn this goal into a practical reality.
— Qing
2021: Expanding plasma proteome coverage to 2,200 proteins
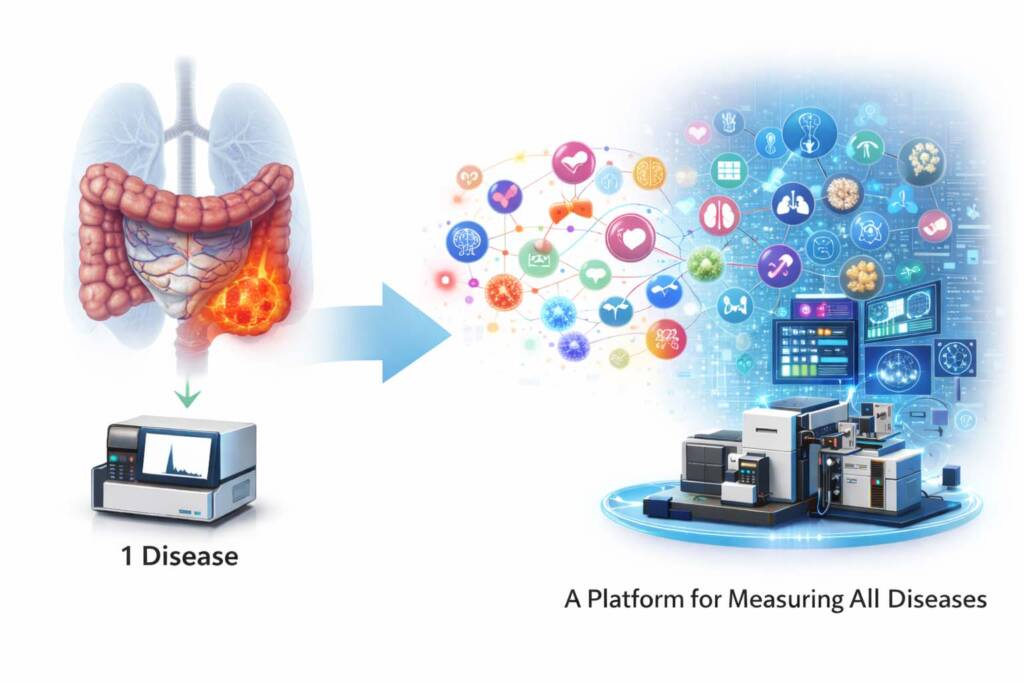
By the end of 2020, this decade-long effort led to another major milestone: the development of workflows capable of detecting approximately 2,200 proteins from plasma samples.
This scale of measurement demonstrated that deep, reproducible coverage of the plasma proteome was becoming technically achievable, bringing the field closer to standardized large-scale proteome measurement. Such capability opens the door to Population-scale proteomic profiling, Early disease detection using blood-based molecular signals, Large-scale biomarker discovery and verification, and Precision therapeutic monitoring
Looking forward: building the measurement layer for precision medicine
Complete Omics is now advancing next-generation proteomics platforms designed to further expand coverage of the plasma proteome while maintaining the quantitative reproducibility required for clinical deployment.
Ten years ago, we demonstrated that mutant proteins could be detected in body fluids. Since then, the question has evolved from whether proteins in plasma can be measured accurately to how deeply and reproducibly we can measure them. Our goal is now to build the analytical infrastructure that makes the human proteome a routine measurement layer for medicine, starting with all human proteins in blood.
— Qing
About Complete Omics—
Complete Omics is building the measurement infrastructure for the human proteome, enabling large-scale, reproducible detection of thousands of proteins from complex biological samples such as plasma. By combining advanced mass spectrometry, engineered workflows, and AI-driven analytics, the company is transforming proteomics from a research discipline into a scalable clinical platform for precision medicine. Complete Omics operates a CLIA-certified, CAP-accredited clinical laboratory, enabling the rapid translation of its technologies into deployable clinical assays. The company’s mission is to make the human proteome measurable at scale, creating a new molecular layer of medicine that powers early disease detection, therapeutic monitoring, and next-generation drug development across every disease.